This work has been funded by the National Science Foundation.
Historically genome size was thought to correlate with organismal complexity, such that larger genomes contain more genes that encode for more complex organisms. However, genome sequencing has shown that genomes are composed of more than just protein-encoding genes. In many cases, just a few genes can be responsible for large phenotypic differences. For example, the transition from teosinte to modern maize involved changes in only five genes.
Others have proposed that genomes have a natural tendency to expand over time and accumulate deleterious mutations unless selection keeps them slim. Because natural selection tends to be more efficient in large populations (in small populations genetic drift is more likely)–so the thinking goes–genome size is a function of population size. Comparisons across widely divergent lineages that span all of life generally support this idea, but over narrower taxonomic scales there is little to no support (Roddy et al. 2021).
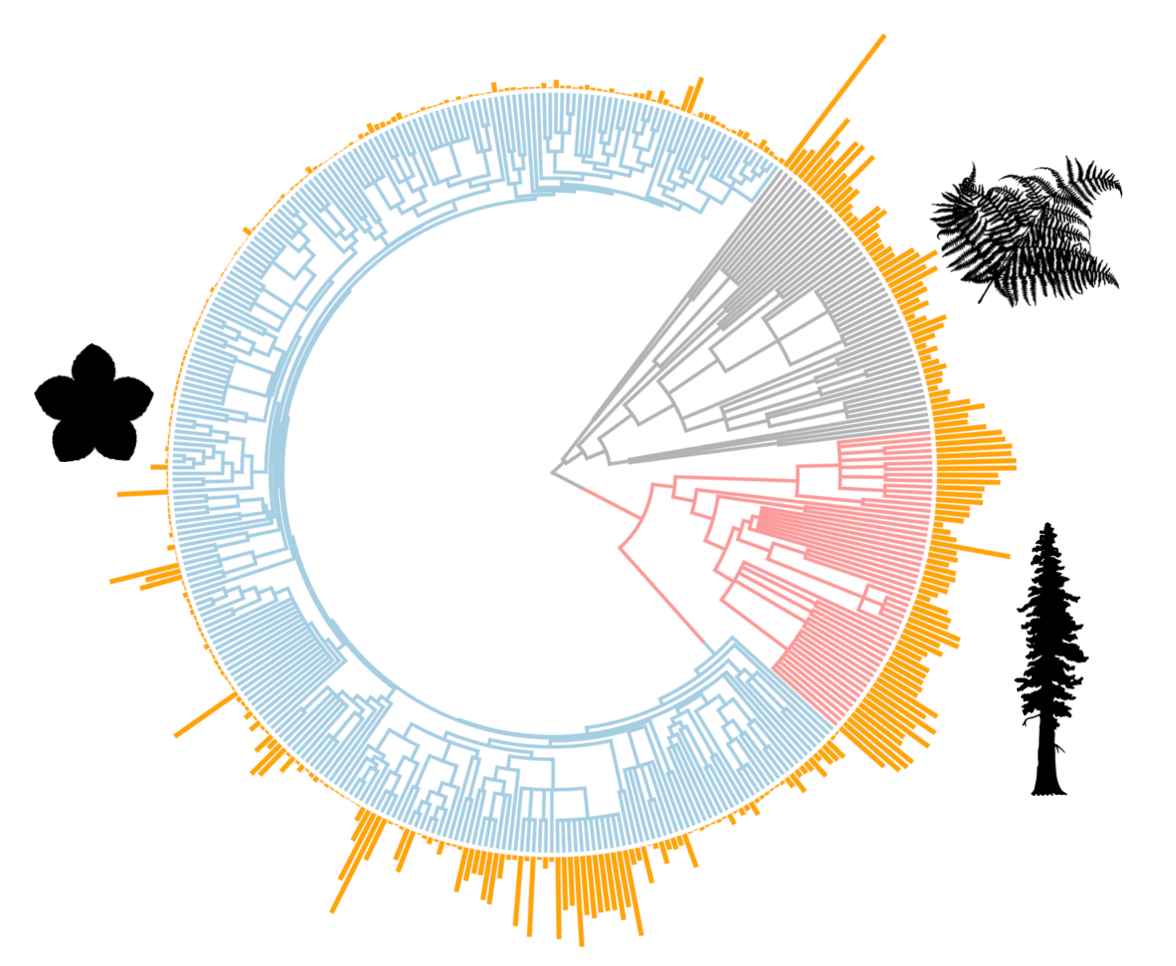

Genome size (pg, orange bars) varies more than 2000-fold across vascular plants (Simonin and Roddy 2018, PLoS Biology).
A parallel line of inquiry has highlighted the nucleotypic effects of genome size variation on numerous plant phenotypes, from the sizes of cells and organs to habitat distributions and phenology. Grime (1998, Annals of Botany) argued that the diverse correlates of genome size should warrant it being considered a key functional trait. Yet, understanding the mechanisms underlying the phenotypic impacts of genome size have remained poorly defined.
We are revisiting ideas about how genome size affects plant phenotypes and, by extension, how selection on plant form or function may act on genome size variation.
At a minimum, a cell must contain its genome. Indeed, the volume of meristematic cells (the totipotent cells that give rise to all other cell types), which are only slightly larger than their nucleus, define the minimum possible cell size. Final cell size, however, can vary dramatically depending on a variety of factors (e.g. cell type, resource availability). Furthermore, because smaller genomes allow for smaller cells, smaller genomes allow for a greater range in final cell size. With a greater variation in cell size may come greater phenotypic plasticity, as smaller genomes allow for cell size to be more fine-tuned to environmental conditions.
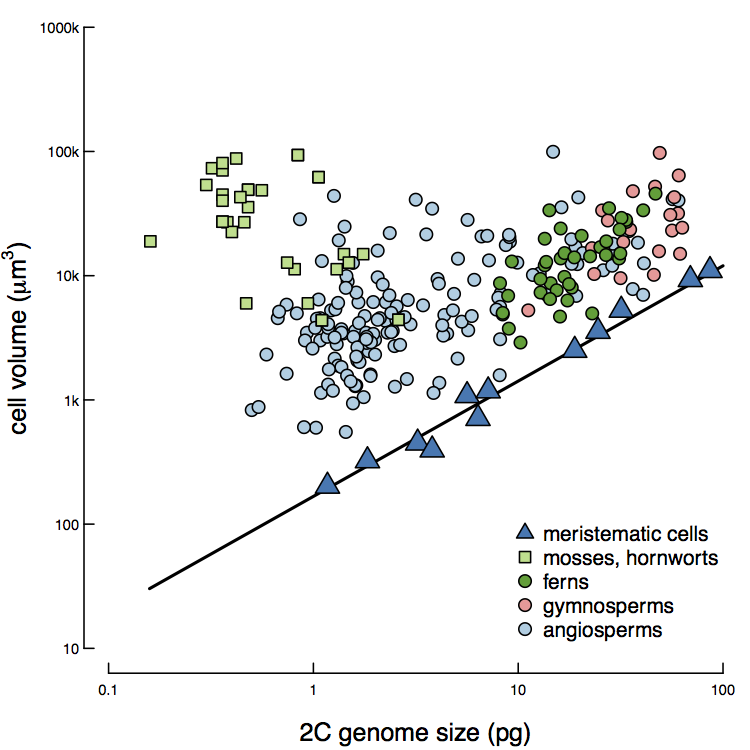
Genome size limits how small a cell can be (Roddy et al., in press).
Cell size–and, thus, genome size–affects metabolism.
First, more smaller cells can be packed into a given space than can larger cells.
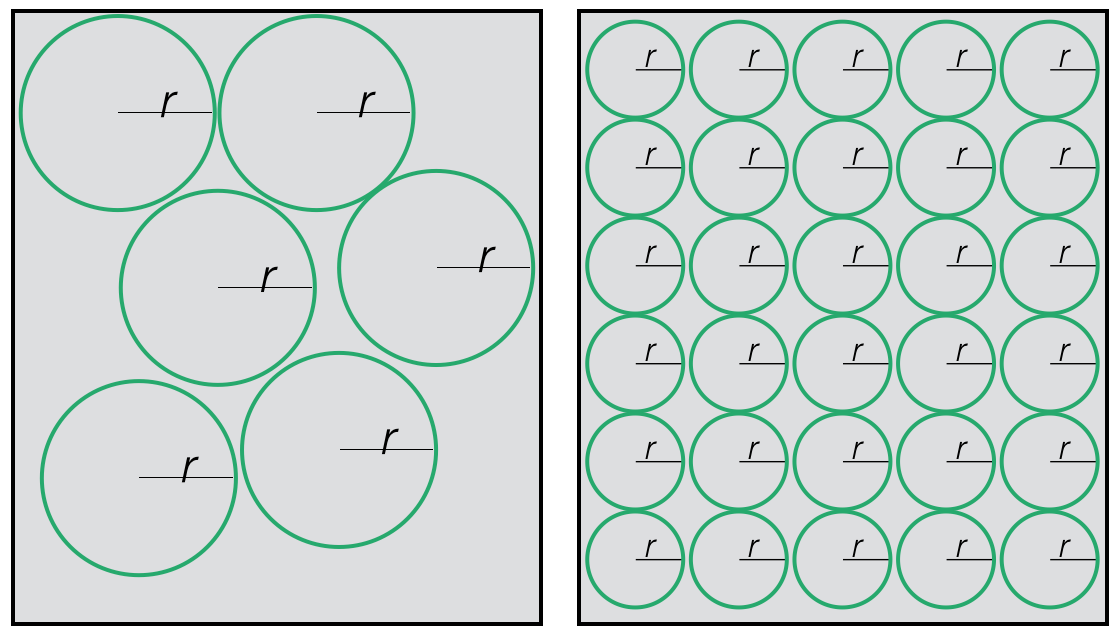
More smaller cells can be packed into a space than larger cells.
Second, smaller cells have a larger ratio of surface area to volume. Because the major limiting factor to metabolism is diffusion, smaller cells can more efficiently move resources into and out of their cells.
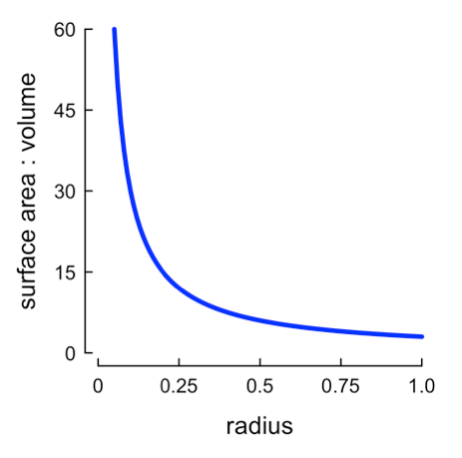
Smaller cells have more surface area for a given volume.
Together, these two points suggest that smaller cells (powered by smaller genomes) have higher rates of metabolism. Despite the effect of genome size being to define the minimum size of a cell, within a cell type, genome size is a major predictor of final cell size across vascular plants. Genome size predicts variation in the sizes and densities of stomata on the leaf surface, through which CO2 and H2O diffuse, and the density of veins in the leaf (Simonin and Roddy 2018, PLoS Biology). These stomatal and vein traits have been shown to influence photosynthetic capacity.
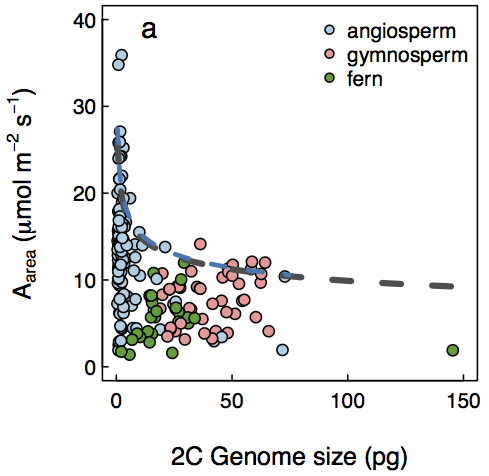
The upper limit (dashed line) to photosynthetic rate is determined by genome size.
Many factors can affect photosynthetic capacity related to the growth environment (nutrient and light availability, etc). When these other factors are not limiting, the maximum attainable photosynthetic rate is limited by cell size and tissue organization.
Cell size variation may have other effects on plant function beyond just limiting maximum metabolic rate. For example, larger cells may confer greater tolerance to certain stresses, such as delaying desiccation or influencing cellular biomechanics (Pacey et al. 2022, Jiang et al. in review)
Related papers
Jiang et al. in review. Mangroves deviate from other angiosperms in their genome size, leaf cell size, and cell packing density relationships.
Borsuk et al. 2022. Structural organization of the spongy mesophyll New Phytologist
Théroux-Rancourt 2021. Maximum CO2 diffusion inside leaves is limited by the scaling of cell size and genome size Proceedings of the Royal Society B
Roddy et al. 2021. Mammals with small populations do not exhibit larger genomes Molecular Biology and Evolution
Roddy et al. 2020. The scaling of genome size and cell size limits maximum rates of photosynthesis with implications for ecological strategies International Journal of Plant Sciences
Simonin and Roddy. 2018. Genome downsizing, physiological novelty, and the global dominance of flowering plants PLOS Biology